An SSA flow model for the Ross Ice Shelf in Antarctica¶
As part of the EISMINT series of intercomparisons, MacAyeal and others [62] successfully validated early-1990s ice shelf numerical models using velocity data for the Ross ice shelf. The data were from the RIGGS survey [135], acquired in the period 1973–1978 and measured at a few hundred locations in a grid across the shelf. Substantial modelling developments followed EISMINT-Ross, including inverse modeling to recover depth-averaged viscosity [136] and parameter-sensitivity studies [137]. Previous PISM versions set up the EISMINT-Ross flow model and performed the diagnostic computation, with RIGGS data for validation.
However, availability of rich new data sets for ice sheet modeling, including the ALBMAP v1 [138] ice sheet geometry, bedrock, and climate data set, and the radar-derived (InSAR) MEaSUREs Antarctica Velocity Map [139], allows us to use more complete, recent, and higher-resolution data for the same basic job. Furthermore one can extend the diagnostic Ross ice shelf calculation both to other ice shelves around Antarctica and to time-evolving (“prognostic”) cases using the eigencalving [99] mechanisms.
The scripts in this subsection are found in directory examples/ross/. In summary, the
script preprocess.py downloads easily-available data and builds a NetCDF input file
for PISM. For the diagnostic computation we document first, the script run_diag.sh (in
subdirectory examples/ross/diagnostic/) runs PISM. The script plot.py shows a
comparison of observations and model results, as in Fig. 42.
Preprocessing input data¶
NSIDC requires registration to download the MEaSUREs InSAR-Based Antarctica Ice Velocity
Map; please register, log in, and get the first, 900 m version
of it (antarctica_ice_velocity_900m.nc) here:
Put this file in examples/ross.
The script preprocess.py then downloads ALBMAP using wget; these two files total
around 350 Mb. Then it uses NCO to cut out the relevant portion of the grid and CDO to
conservatively-interpolate the high-resolution (900 m) velocity data onto the coarser (5
km) geometry grid used in ALBMAP. The script nc2cdo.py from directory util/,
prepares the NetCDF file for the application of CDO, which requires complete projection
information. Run
cd examples/ross/
./preprocess.py
to download ALBMAP and finish the pre-processing.
The NetCDF file Ross_combined.nc produced by preprocess.py contains ice thickness,
bed elevations, surface temperature, net accumulation, as well as latitude and longitude
values. All of these are typical of ice sheet modelling data, both in diagnostic runs and
as needed to initialize and provide boundary conditions for prognostic (evolutionary)
runs; see below for the prognostic case with these data. The _combined file also has
variables u_bc and v_bc for the boundary values used in the (diagnostic
and prognostic) computation of velocity. They are used at all grounded locations and at
ice shelf cells that are immediate neighbors of grounded ice. The variable vel_bc_mask
specifies these locations. Finally the variables u_bc,v_bc, which contain
observed values, are used after the run to compare to the computed interior velocities.
Diagnostic computation of ice shelf velocity¶
The diagnostic velocity computation bootstraps from Ross_combined.nc and performs a
short run; in the \(211\times 211\) grid case we demonstrate below, the key parts of the
PISM command are
pism -i ../Ross_combined.nc -bootstrap -Mx 211 -My 211 -Mz 3 -Lz 3000 -z_spacing equal \
-surface given -stress_balance ssa -energy none -no_mass -yield_stress constant -tauc 1e6 \
-pik -ssa_dirichlet_bc -y 1.0 -ssa_e 0.6 -ssafd_ksp_monitor
The computational grid here is the “native” \(5\) km data grid used in ALBMAP. Regarding the options,
The maximum thickness of the ice is \(2766\) m so we choose a height for the computational box large enough to contain the ice (i.e.
-Lz 3000). Vertical grid resolution is, however, unimportant in this case because we use the SSA stress balance only, and the temperature set at bootstrapping suffices to determine the ice softness; thus the options-Mz 3 -z_spacing equal.Option
-stress_balance ssaselects the SSA stress balance and turns off the SIA stress balance computation, since our goal is to model the ice shelf. It also side-steps a technical issue: PISM uses periodic boundary conditions at domain boundaries and most fields in this setup are not periodic. Turning off SIA avoids operations such as differencing surface elevation across the domain edges. For a more complete solution to this technical issue see section Example: A regional model of the Jakobshavn outlet glacier in Greenland about a regional model using PISM’s “regional mode”pism -regionaland the option-no_model_strip.Option
-y 1.0 -no_mass -energy nonechooses a “diagnostic” run: in absence of geometry evolution and stability restrictions of the energy balance model a one-year-long run will be covered by exactly one time step.Option
-pikis equivalent to-cfbc -part_grid -kill_icebergs -subglin this non-evolving example. Note that-kill_icebergsremoves effectively-detached bits of ice, especially in McMurdo sound area, so that the SSA problem is well-posed for the grounded-ice-sheet-connected ice shelf.Option
-ssa_dirichlet_bcforces the use of fieldsu_bc, v_bc, vel_bc_maskdescribed above. The fieldvel_bc_maskis \(1\) at boundary condition locations, and \(0\) elsewhere. For the prognostic runs below, the ice thickness is also fixed at boundary condition locations, so as to prescribe ice flux as an ice shelf input.Options
-yield_stress constant -tauc 1e6essentially just turn off the grounded-ice evolving yield stress mechanism, which is inactive anyway, and force a high resistance under grounded ice so it does not slide.Option
-ssa_e 0.6is the single tuned parameter; this value gives good correlation between observed and modeled velocity magnitudes.Option
-ssafd_ksp_monitorprovides feedback on the linear solver iterations “underneath” the nonlinear (shear-thinning) SSA solver iteration.
There is no need to type in the above command; just run
cd diagnostic/
./run_diag.sh 2 211 0.6
Note run_diag.sh accepts three arguments: run_diag.sh N Mx E does a run
with N MPI processes, an Mx by Mx grid, and option
-ssa_e E. The choices above give a run which only takes a few seconds, and it
produces output file diag_Mx211.nc.
There are many reasonable choices for the effective softness of an ice shelf, as ice
density, temperature, and the presence of fractures all influence the effective softness.
Using an enhancement factor -ssa_e 0.6 acknowledges that the physical justification
for tuning the ice softness is uncertain. One could instead use the temperature itself or
the ice density[1] as tuning parameters, and these are worthwhile experiments for the
interested PISM user.
The script plot.py takes PISM output such as diag_Mx211.nc to produce
Fig. 42. The run shown in the figure used an enhancement factor of
\(0.6\) as above. The thin black line outlines the floating shelf, which is the actual
modeling domain here. To generate this figure yourself, run
../plot.py diag_Mx211.nc
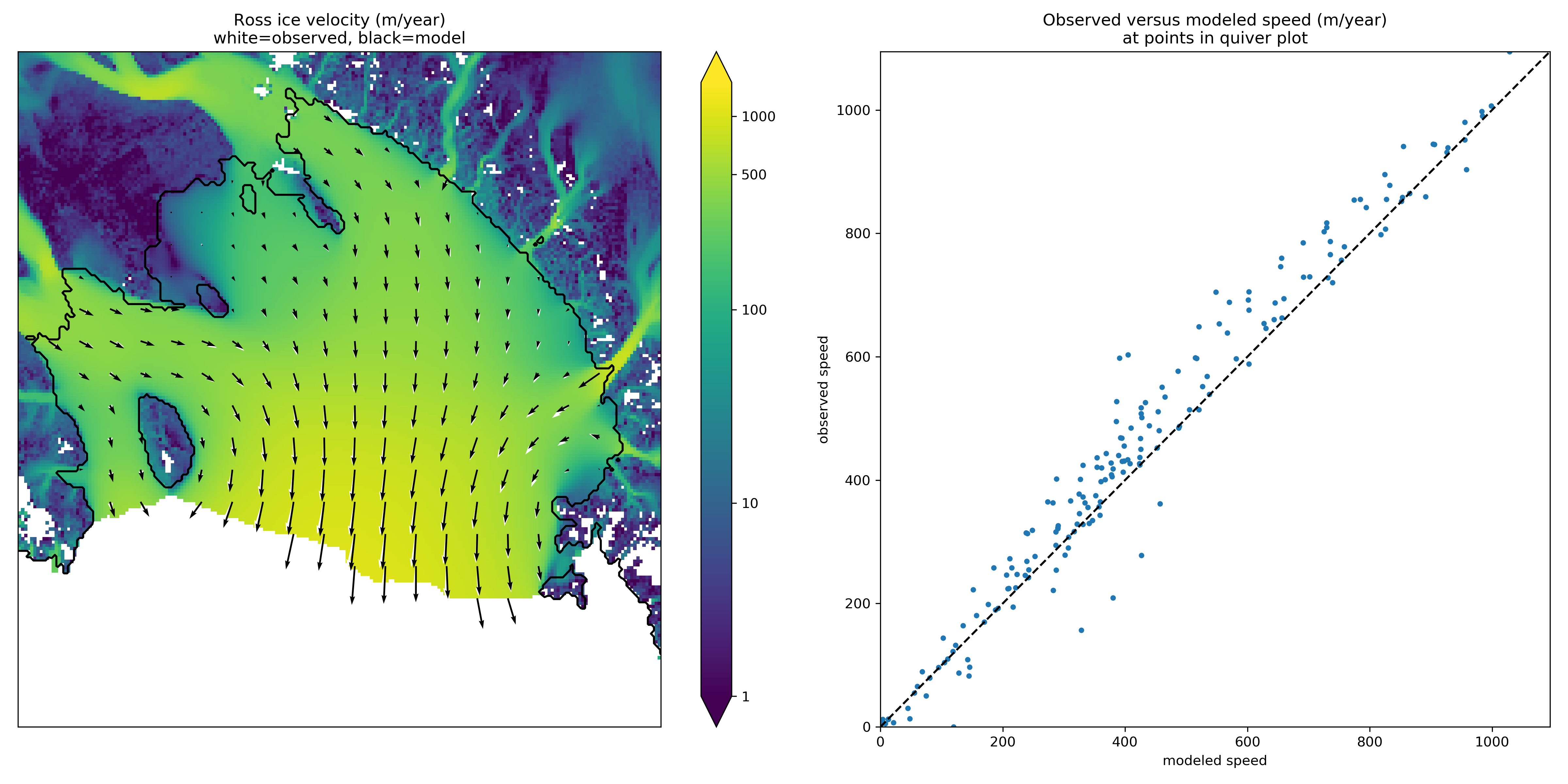
Fig. 42 Left: Color is speed in m/a. Arrows are observed (white) and modeled (black) velocities. Right: Comparison between modeled and observed speeds at points plotted on the left.¶
Extending this example to other ice shelves¶
The SSA diagnostic solution described in this section can be easily applied to other ice shelves in Antarctica, such as the Filchner-Ronne Ice Shelf modeled using PISM in [106], for example.
Simply choose a different rectangular domain, within the area covered by the
whole-Antarctic data-sets used here, at the preprocessing stage. In particular you should
modify the lines “ncks -O -d x1,439,649 -d y1,250,460 ...” (for ALBMAP data) and
“ncks -d x,2200,3700 -d y,3500,4700 ...” (for MEaSUREs velocity data) in the
script examples/ross/preprocess.py.
Prognostic modelling using eigencalving¶
Next we summarize how you can create an evolving-geometry model of the Ross ice shelf with
constant-in-time inflow across the fixed grounding line. See README.md and
run_prog.sh in examples/ross/prognostic/. This example also demonstrates the
-calving eigen_calving model for a moving calving front [99].
Start by running preprocess.py in examples/ross/ as described above. If
you have already done the diagnostic example above, then this stage is complete.
Then change to the prognostic/ directory and run the default example:
cd examples/ross/prognostic/
./run_prog.sh 4 211 0.6 100
This 100 model year run on 4 processes and a 5 km grid took about forty minutes on a 2016
laptop. It starts with a bootstrapping stage which does a -y 1.0 run, which generates
startfile_Mx211.nc. It then re-initializes to start the prognostic run itself. See the
README.md for a bit more on the arguments taken by run_prog.sh and on viewing the
output files.
The PISM command done here is (essentially, and without showing diagnostic output choices)
pism -i startfile_Mx211.nc -surface given -stress_balance ssa \
-yield_stress constant -tauc 1e6 -pik -ssa_dirichlet_bc -ssa_e 0.6 \
-y 100 -o prog_Mx211_yr100.nc -o_size big \
-calving eigen_calving,thickness_calving -eigen_calving_K 1e17 \
-front_retreat_cfl -thickness_calving_threshold 50.0
Several of these options are different from those used in the diagnostic case. First,
while the command -pik is the same as before, now each part of its expansion, namely
-cfbc -part_grid -kill_icebergs -subgl, is important. As the calving front evolves
(i.e. regardless of the calving law choices), option -part_grid moves the calving
front by one grid cell only when the cell is full of the ice flowing into it; see
[98]. The option -kill_icebergs is essential to maintain well-posedness
of the SSA velocity problem at each time step [17]. See section
PIK options for marine ice sheets.
Option combination
-calving eigen_calving,thickness_calving -eigen_calving_K 1e17 \
-front_retreat_cfl -thickness_calving_threshold 50.0
specifies that ice at the calving front will be removed if either a criterion on the
product of principal stresses is satisfied [99], namely eigen_calving
with the given constant \(K\), or if the ice thickness goes below the given threshold of 50
meters. See section Calving and front retreat.
Footnotes
| Previous | Up | Next |
